What is Decision Tree algorithm— overview and an example
About the Author in his words: Vishal R is interested in Machine Learning and it's applications. You can follow the author on his medium or twitter. The article was originally published on towardsdatascience
What are Decision Trees?
A decision tree is a flowchart-like structure in which each internal node represents a “test” on an attribute (e.g. whether a coin flip comes up heads or tails), each branch represents the outcome of the test, and each leaf node represents a class label (decision taken after computing all attributes).In simpler terms, a decision tree checks if an attribute or a set of attributes satisfy a condition and based on the result of the check, the subsequent checks are performed. The tree splits the data into different parts based these checks.
Implementation : Importing the necessary libraries
import numpy as np
from sklearn.preprocessing import StandardScaler
import tflearn.data_utils as du
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import train_test_split
import seaborn as sns
from sklearn.metrics import confusion_matrix
from sklearn.externals.six import StringIO
from IPython.display import Image
from sklearn.tree import export_graphviz
import pydotplus
Reading the dataset
data = pd.read_csv('../input/column_3C_weka.csv')The dataset used here is the Biomechanical features of orthopedic patients
What is correlation?
Correlation is a statistical term which in common usage refers to how close two variables are to having a linear relationship with each other.
For example, two variable which are linearly dependent (say, x and y which depend on each other as x = 2y) will have a higher correlation than two variables which are non-linearly dependent (say, u and v which depend on each other as u = sqr(v))
Visualizing the correlation
# Calculating the correlation matrix
corr = data.corr()
# Generating a heatmap
sns.heatmap(corr,xticklabels=corr.columns, yticklabels=corr.columns)

sns.pairplot(data)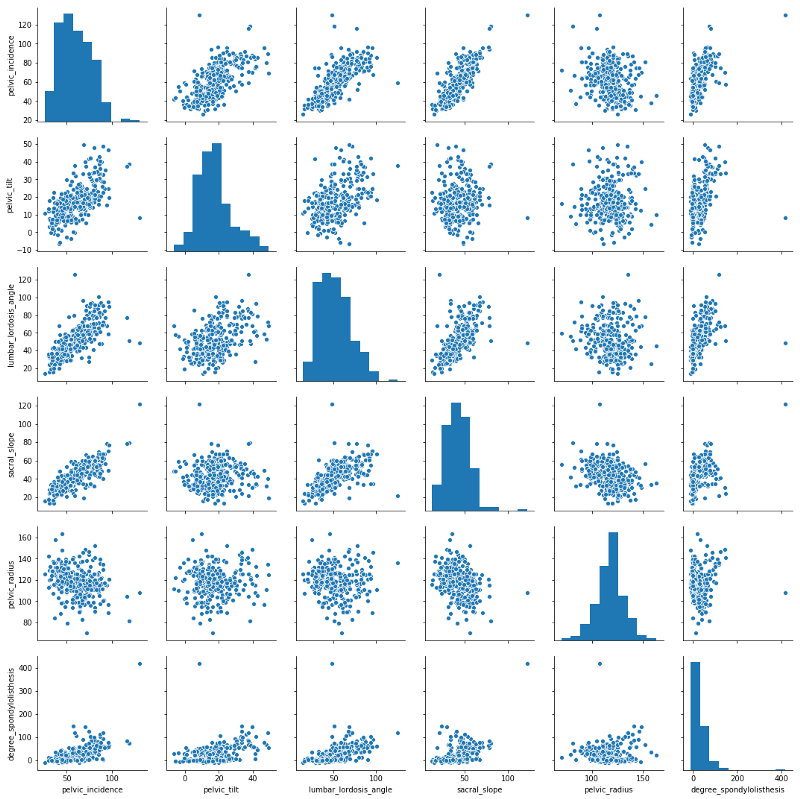
Splitting the dataset into independent (x) and dependent (y) variables
The train data to train the model and the test data to validate the model’s performance
x_train , x_test, y_train, y_test = train_test_split(x, y, test_size = 0.25, random_state = 0)
Scaling the independent variables
This question on stackoverflow has responses which gives a brief explanation on why scaling is necessary and how it can affect the modelsc = StandardScaler()
x_train = sc.fit_transform(x_train)
x_test = sc.transform(x_test)
Building the Decision tree
The criterion here is entropy. The criterion parameter determines the function to measure the quality of a split. When the entropy is used as a criterion, each split tries to reduce the randomness in that part of the data.
There are lot of parameters in the Decision Tree Class that you can tweak to improve your results. There we take a peak into the max_depth parameter.
The max_dept the determines how deep a tree can go. The affect of this parameter on the model will be discusses later in this article
classifier = DecisionTreeClassifier(criterion = 'entropy', max_depth = 4)
classifier.fit(x_train, y_train)
Making the prediction on the test data
y_pred = classifier.predict(x_test)
What is a confusion matrix?
A confusion matrix is a technique for summarizing the performance of a classification algorithm. Classification accuracy alone can be misleading if you have an unequal number of observations in each class or if you have more than two classes in your dataset. Calculating a confusion matrix can give you a better idea of what your classification model is getting right and what types of errors it is making.
cm = confusion_matrix(y_test, y_pred)
accuracy = sum(cm[i][i] for i in range(3)) / y_test.shape[0]
print("accuracy = " + str(accuracy))
Visualizing the Decision Tree
dot_data = StringIO()
export_graphviz(classifier, out_file=dot_data,
filled=True, rounded=True,
special_characters=True)
graph = pydotplus.graph_from_dot_data(dot_data.getvalue())
Image(graph.create_png())
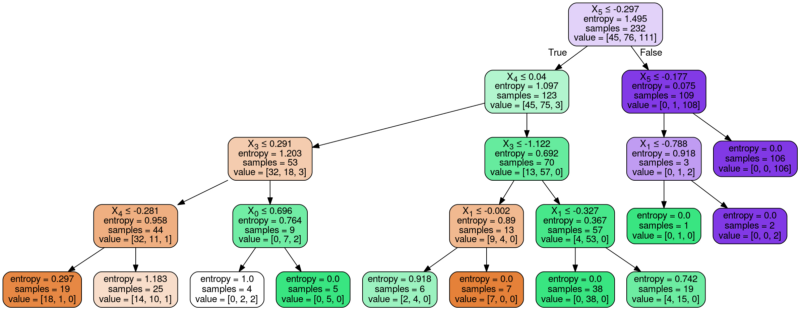
Building a model without the max_depth parameter
classifier2 = DecisionTreeClassifier(criterion = 'entropy')
classifier2.fit(x_train, y_train)
y_pred2 = classifier2.predict(x_test)
cm2 = confusion_matrix(y_test, y_pred2)
accuracy2 = sum(cm2[i][i] for i in range(3)) / y_test.shape[0]
print("accuracy = " + str(accuracy2))
Visualizing the decision tree without the max_depth parameter
dot_data = StringIO()
export_graphviz(classifier2, out_file=dot_data,
filled=True, rounded=True,
special_characters=True)
graph = pydotplus.graph_from_dot_data(dot_data.getvalue())
Image(graph.create_png())
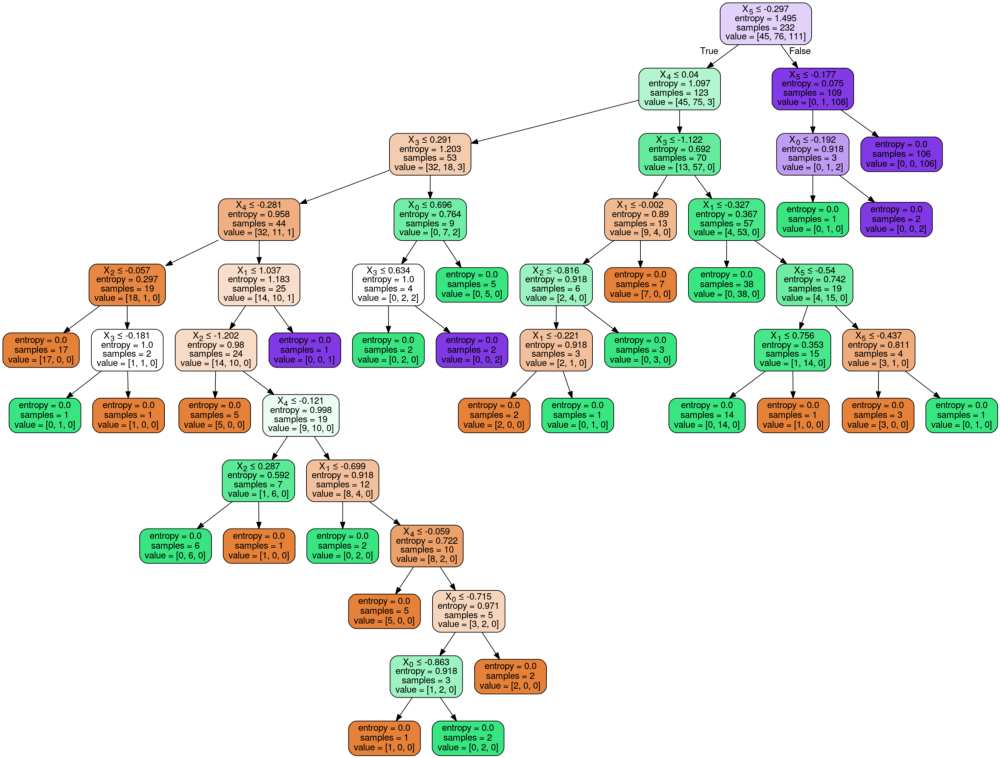
Now, consider the leaf nodes (terminal nodes) of the tree with and without the max_depth parameter. You will notice that the entropy all the terminal nodes are zero in the tree without the max_depth parameter and non zero in three with that parameter. This is because when the parameter is not mentioned, the split recursively takes place till the terminal node has an entropy of zero.
To get this article as an iPython Notebook, click here
victor
posted on 02 Oct 18Enjoy great content like this and a lot more !
Signup for a free account to write a post / comment / upvote posts. Its simple and takes less than 5 seconds

Post Comment